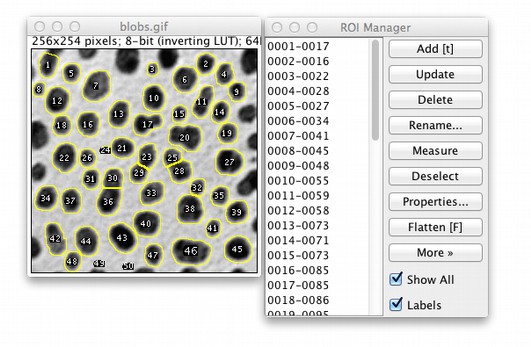
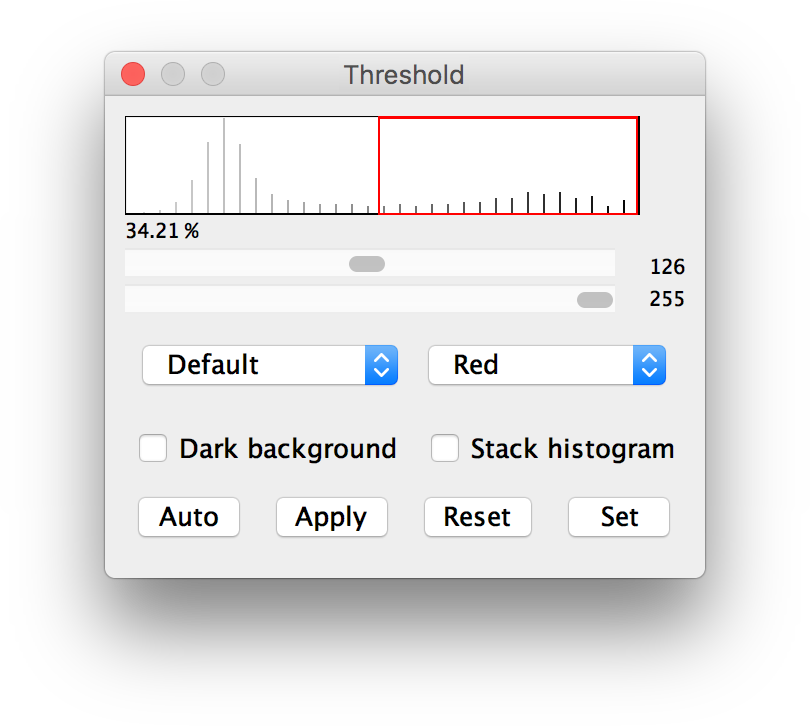
Obtained pore information from the graphical process using the ImageJ is very subjective and uncertain. I feel this method is quite subjective, likely to create great variabilities between images. ASHOK (ashok dewangan) April 6, 2016, 1:04pm 1. ROIs) to cover all the GFP expressed areas in the original image, measure the mean value (3), then subtract (2) from (3). Now comes the question – how do I create ROIs from the thresholded image to cover all black areas (magic wand only highlight individual white areas)? Or is there a way to do the “mapping”? Then, if possible, I could “map” the black areas to my original image to measure the mean value from the “proper” protein expressed area (let’s assume that the autofluorescence is comparable between 2 groups). The white areas then have value of 0 and black areas 255. Threshold the image using range of 0 to value (2) as measured above to get the Thresholded Image.Check Interpolate to use bilinear interpolation. Check Constrain Aspect Ratio to create an image with the specified width and have ImageJ adjust the height to maintain the original aspect ratio. Scales the active image or selection to a specified width and height in pixels.

Open ImageJ and record your processing steps: Plugins -> Macro -> Record: Image -> type -> 8-bit. Easy to achieve, and seems reasonably logical. Click on Set to enter new threshold levels into a dialog box. A note for myself concerning batch processing egg-counting in ImageJ. Measure the mean value (1) within this ROI, then measure the background mean value (2) by using a set of small ROIs placed in those dark areas within the above polygonal ROI.
